CRISPR Knockin Stable Cell Line
VectorBuilder는 원하는 게놈 타겟 부위에 GOI를 knockin한 안정적인 세포주를 제작할 수 있습니다. 안정적인 knockin은 CRISPR 기반 방법을 통해 달성되며, 타겟 부위 특이적 gRNA/Cas9 RNP complex는 HDR (homology-directed repair)을 위한 주형을 운반하는 donor vector와 함께 타겟 세포에 도입됩니다. VectorBuilder의 독점 기술을 통해 매우 효율적인 유전자 전달 및 HDR을 달성할 수 있고, homozygous knockin 제작에 대한 신속한 소요시간과 높은 성공률을 달성할 수 있습니다.
중점 사항
- Supreme integration efficiency: 독자적인 donor 디자인 및 유전자 전달 방법을 통해 대부분의 세포는 30-50%의 HDR 비율을 나타냅니다.
- Large fragment knockin: 최대 1 kb의 단편을 ESC 및 iPSC를 포함한 다양한 세포에 효율적으로 도입할 수 있습니다.
- Rapid turnaround: monoclonal knockin 세포는 벡터 디자인부터 시작하여 빠르면 16주 만에 제작할 수 있습니다.
서비스 상세
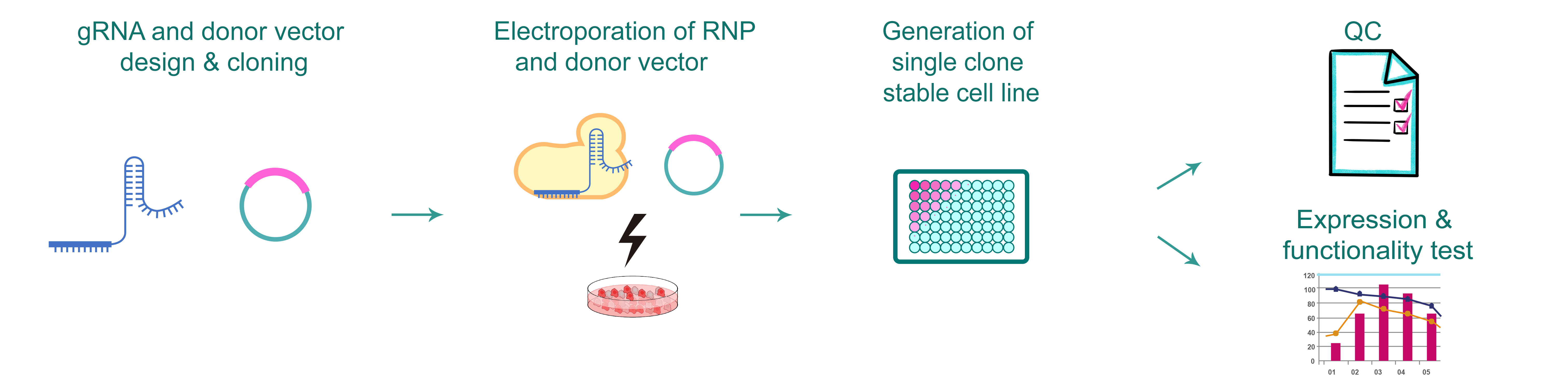
Figure 1. Typical workflow of generating gene knockin cells by CRISPR.
가격 및 소요시간
| Service Type | Deliverable | Price (USD)* | Turnaround |
|---|---|---|---|
| Knockin (fragment<3 kb) | One heterozygous single clone (>106 cells/vial, 2 vials) | From $8,999 | 14-20 weeks |
| One homozygous single clone (>106 cells/vial, 2 vials) | Please inquire | ||
| Knockin (fragment>3 kb) | One heterozygous single clone (>106 cells/vial, 2 vials) | From $11,999 | 18-25 weeks |
| One homozygous single clone (>106 cells/vial, 2 vials) | Please inquire | ||
* Additional charge will apply for extra single clones or vials.
QC assays
| Assay | Methods |
|---|---|
| Knockin validation (default) | Genotyping PCR, Sanger sequencing |
| Expression test (add-on) | RT-qPCR, WB, IF, FACS |
| Off-target analysis (add-on) | NGS, PCR, Sanger sequencing |
| Chromosome analysis (add-on) | Karyotyping |
| Sterility (default) | PCR for mycoplasma detection, bioburden test for sterility detection |
다운스트림 서비스
VectorBuilder는 proliferation, apoptosis, migration, viability, cytotoxicity 등에 대한 분석을 포함하여 제작된 세포주의 다양한 phenotype 평가 및 기능 검증을 수행할 수 있습니다.
사례 연구
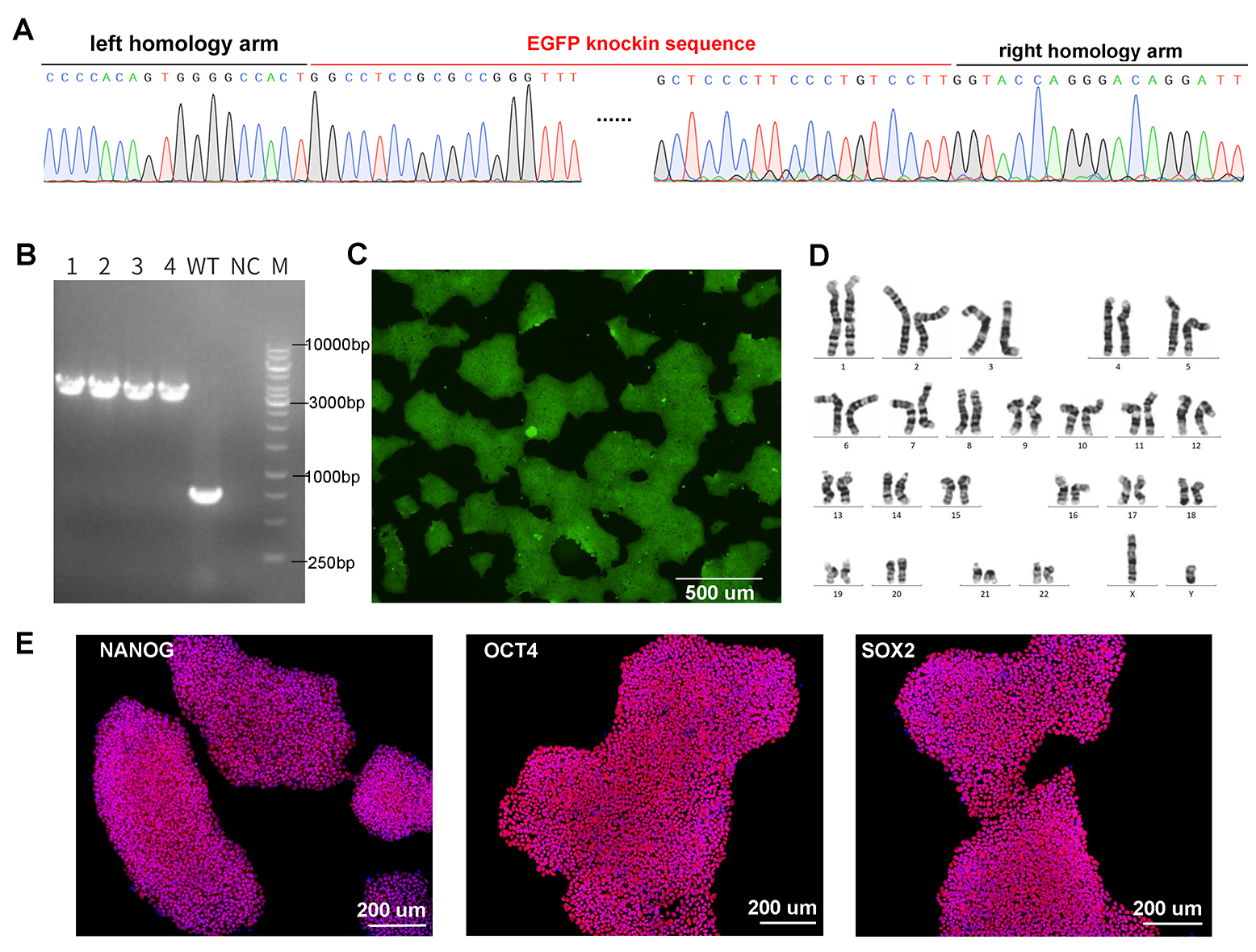
Figure 2. CRISPR-mediated gene knockin in iPSCs. Knockin of UBC-driven EGFP (2432 bp) into iPSCs was achieved by electroporation of Cas9/gRNA RNP complex and donor vector. (A) Confirmation of EGFP knockin at target site by Sanger sequencing. (B) Genotyping PCR of four single clones with homozygous knockin. The WT locus is 762 bp and the locus with EGFP knockin is 3194 bp. (C) EGFP fluorescence in knockin cells by microscopy. (D) Karyotyping results. (E) Expression of pluripotency markers NANOG, OCT4, and SOX2 in EGFP knockin iPSCs by immunofluorescence.
주문 방법
FAQ
Gene knockout relies on non-homologous end joining (NHEJ) mediated DNA repair of double strand breaks (DSBs) caused by CRISPR which is error-prone. As a result, mutations such as frameshift or fragment deletion (when using dual-gRNA) happen at a certain frequency which cause loss-of-function of the target gene. Differently, gene knockin depends on precise insertion of donor fragment at target site by HDR which is much less efficient than NHEJ and is limited to dividing cells. Additionally, HDR efficiencies vary by cell type. Altogether, gene knockin is technically more challenging than gene knockout.
The primary consideration for the format of your template is size. If introducing a small insert (<10 bp), then ssODNs are typically the best option because of its enhanced efficiency of HDR. Larger templates must be introduced with dsDNA in either circular plasmid form (higher cell tolerance but lower HDR efficiency) or linear form (higher HDR efficiency but higher cell toxicity).
Following transfection and antibiotic drug selection, a pool of positively transfected cells is established. Cells are counted, e.g. with a hemocytometer, to determine cell number and concentration. Cells then go through serial dilution (limiting dilution) and are cultured in 96-well plates at a concentration of <1 cell per well. These individual viable clones are expanded and genotyped to confirm the desired genetic modification.



