Vector Systems
Worm Regular Plasmid Gene Expression Vector
Overview
Microinjection of DNA into the distal arm of the gonad of C. elegans is a popular tool for gene delivery. In this technique, regular plasmids are injected and incorporated into germ cell nuclei. The injected DNA tends to form an extrachromosomal array and can show drift of expression over generations. The choice of tissue specific and inducible promoters can be a useful tool in designing a vector for your C. elegans experiment.
For further information about this vector system, please refer to the papers below.
| References | Topic |
|---|---|
| EMBO J. 10:3959 (1991) | DNA transformation in C. elegans |
| Sci Rep. 9:9192 (2019) | Promoter-proximal introns increases gene expression levels in C. elegans. |
Highlights
Our vector is optimized for inducing high gene expression in C. elegans by incorporating a synthetic promoter-proximal intron in the backbone.
Advantages
Technical simplicity: Delivery of plasmid vectors into cells by microinjection is technically straightforward and far easier than virus-based vectors which requires the packaging of live virus.
Very large cargo space: Our vector can accommodate ~30 kb of total DNA. The plasmid backbone only occupies ~2.25 kb, leaving plenty of room to accommodate the user’s sequence of interest.
High-level expression: A synthetic promoter-proximal intron is incorporated into the vector backbone. This leads to very high expression levels of the genes carried on the vector.
Disadvantages
Non-integration of vector DNA: Regular plasmid vectors mostly remain as episomal DNA, without genome integration. However, plasmid DNA can integrate permanently into the host genome at a low frequency. If a drug resistance cassette is incorporated into the plasmid, cells that have stably integrated the plasmid can be derived by drug selection after extended culture.
Limited cell type range: The efficiency of regular plasmid delivery can vary greatly from cell type to cell type.
Non-uniformity of gene delivery: Although high copy numbers of transgenes can be achieved, this can be non-uniform. Some cells may carry many copies while others may carry very few, or none.
Key components
Promoter: The promoter driving your gene of interest is placed here.
Synthetic intron: This is placed in proximal to promoter to promote gene expression.
Kozak: Kozak consensus sequence. This is placed in front of the start codon of the ORF of interest to facilitate translation initiation in eukaryotes.
ORF: The open reading frame of your gene of interest is placed here.
3' UTR+polyA: Allows transcription termination and polyadenylation of mRNA transcribed by RNA polymerase II. It functions in somatic and germ cells.
Ampicillin: Ampicillin resistance gene. It allows the plasmid to be maintained by ampicillin selection in E. coli.
pUC ori: pUC origin of replication. Plasmids carrying this origin exist in high copy numbers in E. coli.
Experimental validation
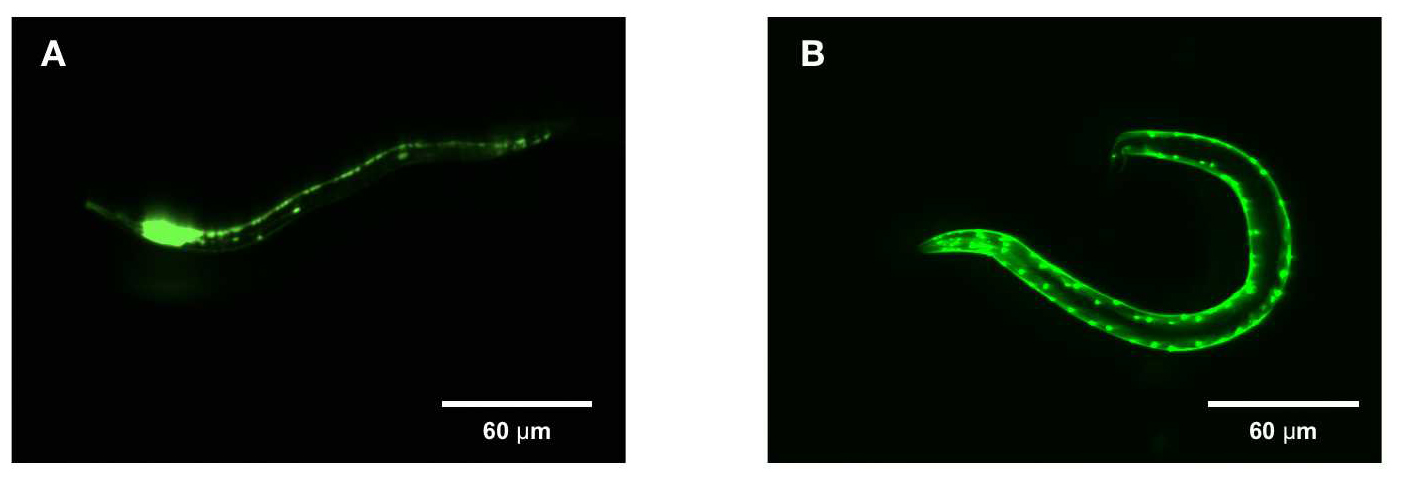
Tissue-specific EGFP expression in C. elegans.
Parental C. elegans (P0, N2 strain) were microinjected with EGFP expression plasmids (32 ng) and DNA ladder (256 ng) into the distal arm of the gonad. The rab-3 and myo-2 promoters respectively drive (A) neuron and (B) muscle-specific EGFP expression in their adult progeny (P1) 3 days post-hatching. Heads are oriented to the left. Scale bar=60 um.



